December 2019 Release
GlyTouCan v3.1.0 (December 12, 2019)
- Registration system updated.
- Glycans can be registered using GlycoCT format.
- The GlycoCT conversion to WURCS batch process is running.
- Update text input form for registration and the confirmation page when registering has been abolished.
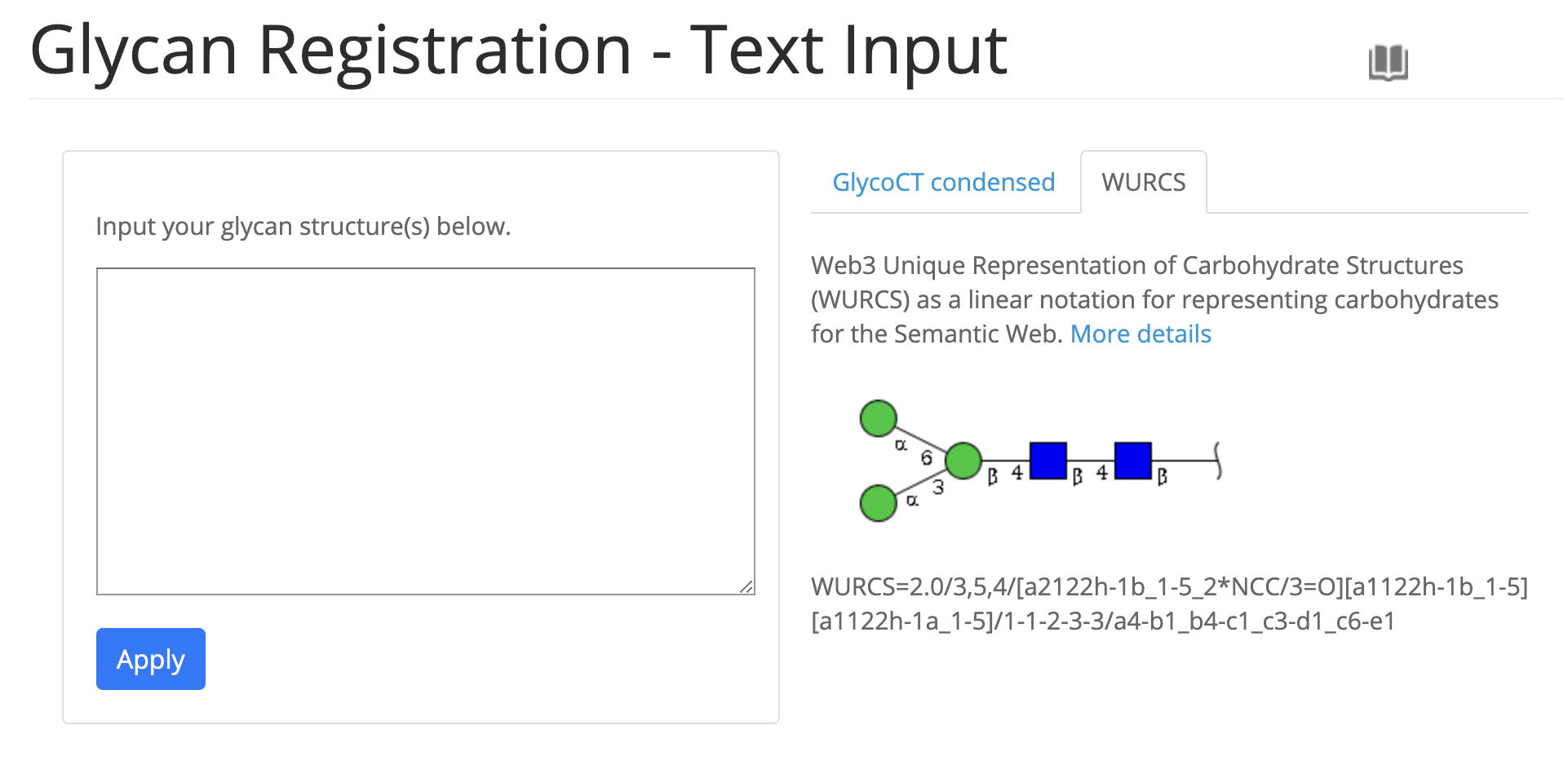
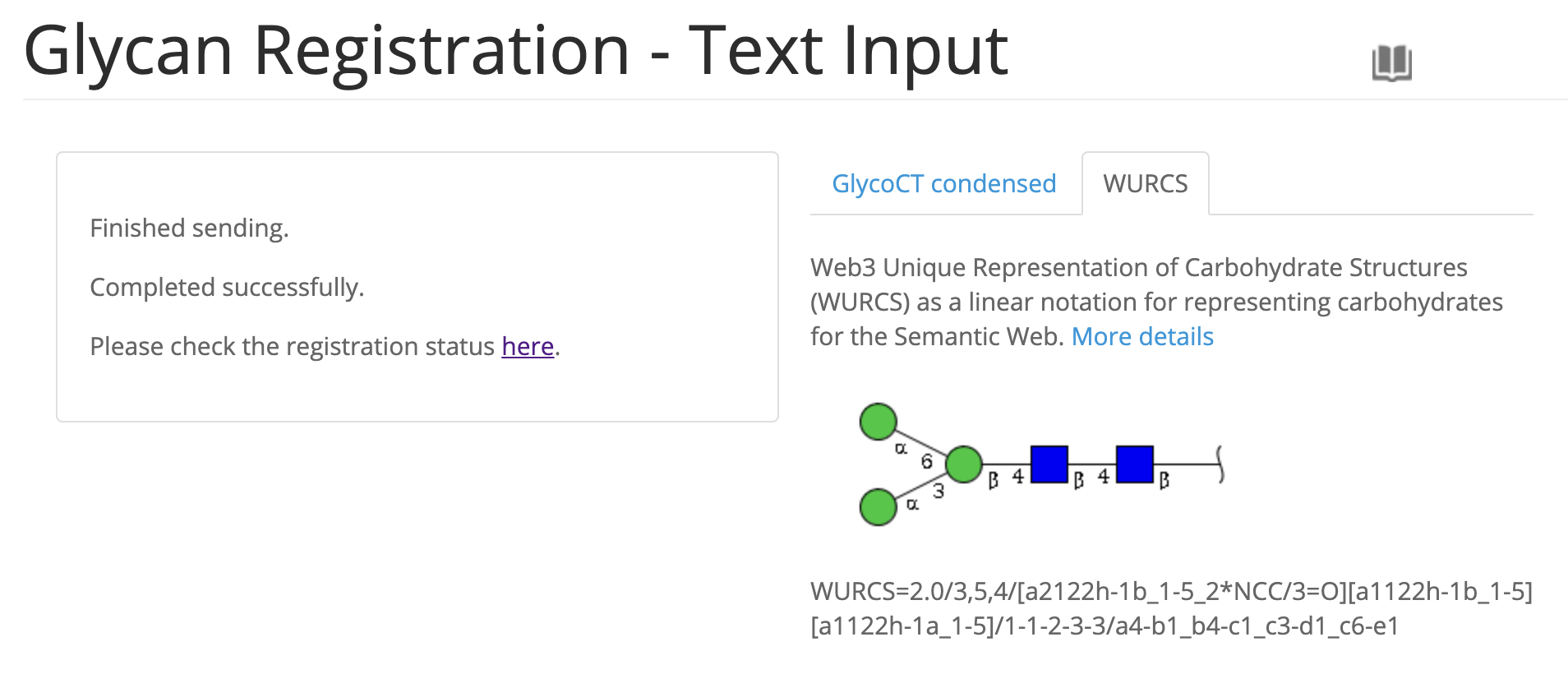


- Improved submissions page.
- pagination, line select, show the total submissions and accession numbers as registered.
August 2019 Release
New Registration Flow and Backend Framework
This past week, a major release and infrastructure change was made to the GlyTouCan system. Here is a brief rundown of features and then a more detailed explanation of how these changes will impact users.
It has been a while since the last post. One reason is that the initial GlyTouCan project officially ended, while planning and expansion of the system was included in the new GlyCosmos Project.
Main Features
- New structure registration process: Any structure format or nomenclature accepted
- More transparency of how inputted structures are detected, converted, validated, etc
- Background batch-processing framework introduced
- Audited data conversion/modification/validation
- Image background processing
- New server hardware
- New Entries page which shows all previous registrations
- New UI methodology
- Improved Continuous Integration (CI) Development cycle (for developer’s sanity and stability)
Questions Answered
This release answers a lot of questions that we were receiving. In particular the following:
- “Why can’t I input my structure in IUPAC/GlycoCT/KCF/… format?”
- “I have doubts about this structure, how was it generated?”
- “The image is wrong!”
New Registration Flow
The above questions pointed at an initial design flaw of the repository. The policy to assign an Accession Number was the following: “If the structure could be converted into WURCS, then it was assigned an Accession Number”. The problem with this policy is that it assumes all structures input can be converted into WURCS. If it could not, then an error was returned. Also, since the registration system solely was an API server, the many steps required (detection/conversion/validation) were executed once the submit button was pressed. Error management handling was not very user-friendly. These errors were not recorded or logged, which made it difficult to support. This showed how the system was not future-proof, and so a major architecture shift was required.
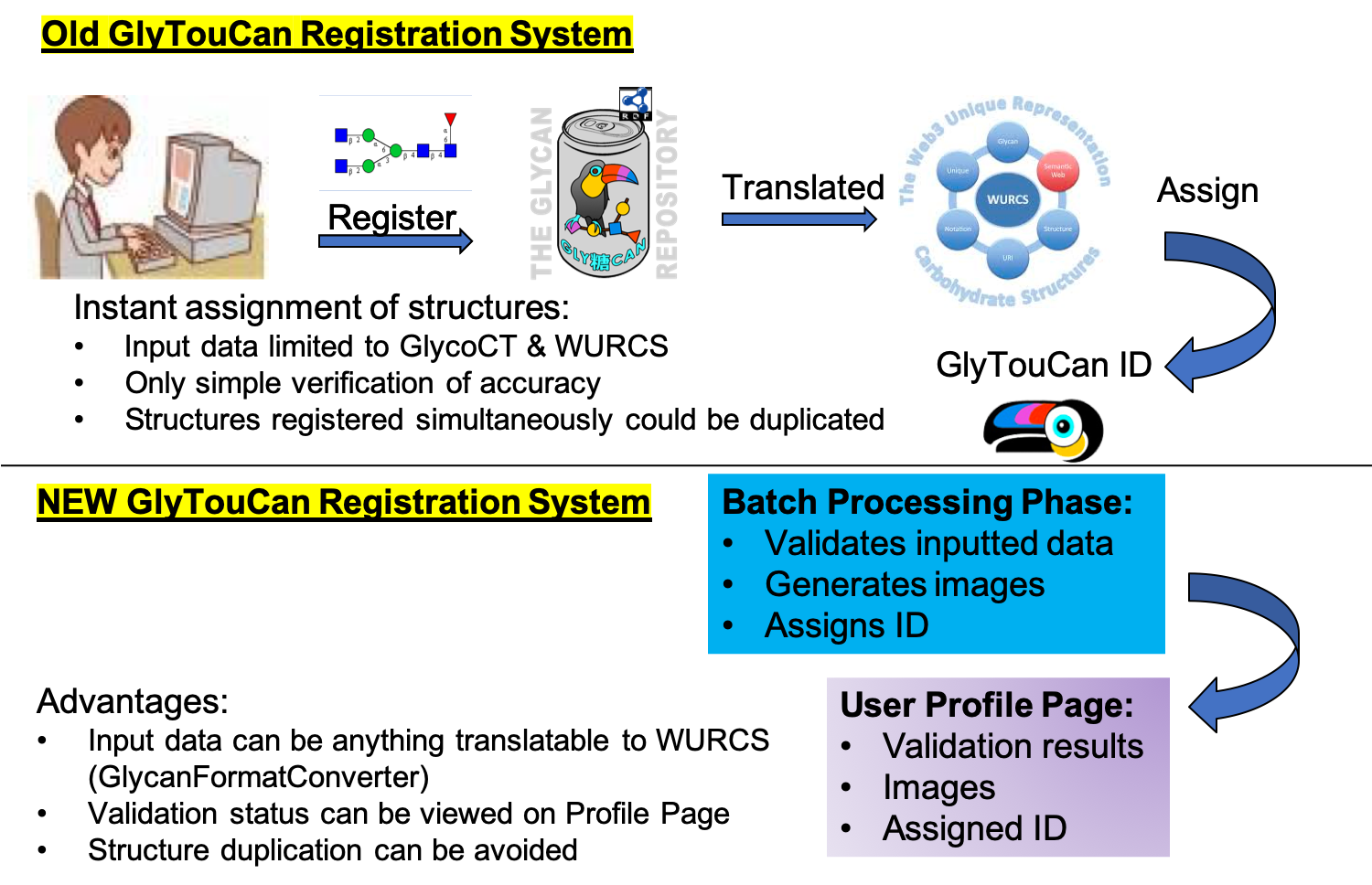
This release introduces a new registration flow. The new policy is the following: “Any structure sequence format can be input”. Once “pre-registered”, the submission will be given a reference tag. This tag can be used to lookup the structure at a later time. In case this tag is lost, all previously submitted structures are displayed in the new personalized Entries page.
The steps described above (detection/conversion/validation) are now split up into multiple server-side, backend batch processes. These are used to work on the submitted structure, such as detecting the nomenclature/format, converting it to WURCS(if necessary), validating it, assigning an Accession Number, or generating the image etc.
The status of these batch processes will be visible through the Entries page as well as the detailed entry page of each structure submitted. As a general policy all batch process information will be transparent to the user. We will be constantly updating the UI to show as much information as possible returned by these batch processes.
This will cause some delay between initial registration and Accession Number assignment. However it will help clarify what exactly are the issues with the structure, if there are any, for very specific steps in the flow. By removing all complex logic libraries from the website/API, more stability is guaranteed as there are less points of failure.
Detailed Registration Example
The following is a simple example of a structure registration, with detailed explanation of the new flow.
- Registration of WURCS structure
- Format Detection
- WURCS Validation
- Accession Number Generation
The first batch process is a simple “structure format detection” batch. This will detect the structure as WURCS and add a new RDF triple indicating that the structure has a format of “wurcs”. It is meant to organize structures into specific formats so that it will be more obvious what batch process needs to run next.
The next batch process contains logic to search only for structures labeled as “wurcs”, and validates them. A specific WURCS library is used which reads the structure in and outputs warnings or error messages. These warnings and error messages are now stored into the RDF and linked to the structure. If there are error messages, it is not considered validated. Since the warnings and error messages are now stored, the reason why the validation failed can now be displayed to the user on the Entries page.
The next process searches for validated structures, and contains logic to see if the structure was already registered or not. If previously registered, the structure will be assigned the previous Accession Number. Otherwise it will generate a new Accession Number for the new structure. Once the Accession Number is assigned, the previous GlyTouCan system will continue to run as before.
New Image Batch Processing
A new image generation batch process will convert the input structures into multiple notations and formats. All of the different combinations of SNFG, CFG, IUPAC and SVG, PNG will be generated and stored locally. The main problem why there were weird images appearing before was due to generating the image in real-time for every user request. Since this CPU intensive and non-multi-threaded process is now put off into the background batch process, each user request will be a simple read of the correct image raw data.
Known issues
The GlycoCT conversion to WURCS batch process is in development. It will be introduced within the next few weeks.
The website still does some conversion automatically. This will be removed once the GlycoCT batch is complete.
One of the major changes with this process is that the Accession Number is not received in real-time after submission. The Website, client library, and CLI interface were all slightly modified to account for this. As a simple measure to prevent duplications, an exact search of the input sequence is done to check if it is registered already or not. If so, the accession number is returned, otherwise the reference tag is returned.
Future Work
Since the Batch process is based on a framework, an API to this framework can be created to give user’s direct access to these batch process timings and even direct execution requests.
More work on the user interface. Simple things such as pagination were deprioritized until after the release. These will be hotfixed as they will not impact underlying RDF data. Since we are utilizing a new more modern standardized method of GUI written in javascript, along with a CI development system, it allows us to quickly release updates in seconds.
The new architecture allows us to split up the vast amount of work we had done on the website, and focuses more on discrete logical steps to process each strucure. This allows for a simple way to develop and expand the RDF data in an organized fashion, while keeping the user interface completely stable. Please contact us directly if there are any issues!